🎯
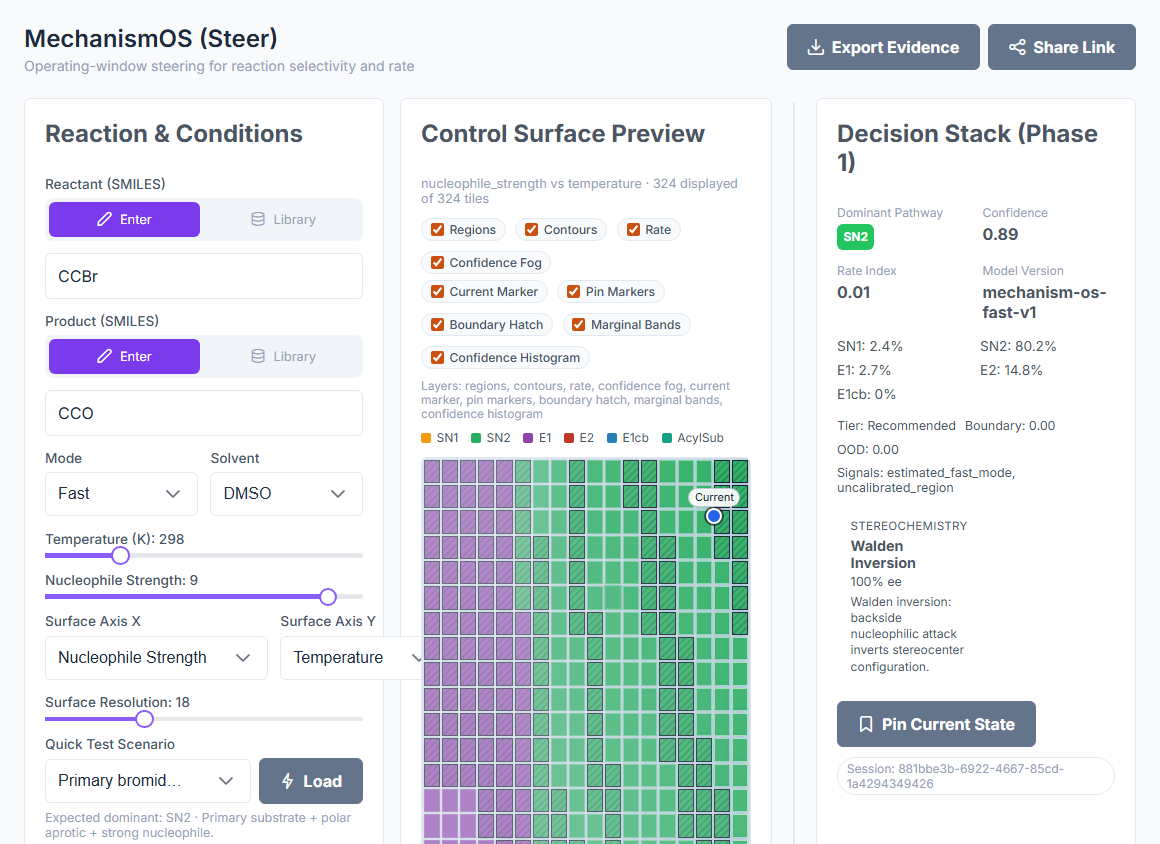
5-way classification
SN1, SN2, E1, E2, E1cb — all five mechanisms from a single call, with Boltzmann-weighted competition between them.
⚡
DFT-grade barriers
7.4 kJ/mol MAE on activation energies — inside the DFT accuracy window — in ~1 ms instead of hours.
🌡️
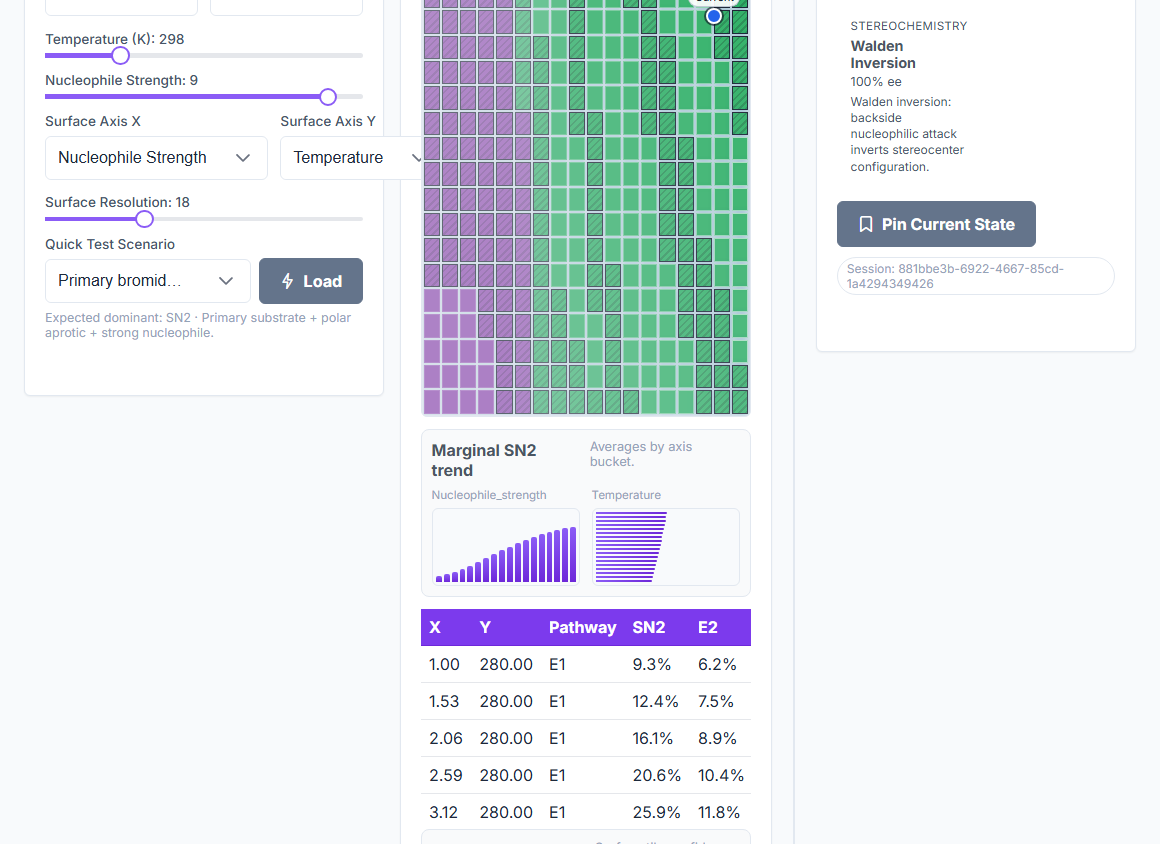
Temperature crossovers
Predicts SN1→E1 and SN2→E2 crossovers as temperature changes, including the crossover temperature itself.
🔀
Rearrangement detection
Hydride shifts, methyl shifts, and ring expansions flagged automatically under SN1 / E1 conditions via graph traversal.
🔗
Neighbouring-group participation
Anchimeric assistance from S, N, and phenyl groups detected with rate-enhancement estimates (episulfonium, aziridinium, phenonium).
💧
16 solvents
Water, ethanol, DMSO, DMF, acetone, TFE, t-BuOH, methanol, THF — protic, aprotic, and non-polar solvents all parameterised.
🧬
Full interpretability
Every kJ/mol of the barrier traces back to a physical factor — nucleophilicity, steric strain, solvent stabilisation, leaving-group ability. Not a black box.
🔬
Wide substrate scope
7 substrate classes (methyl, primary, secondary, tertiary, allylic, benzylic, vinyl) × 7 leaving groups (I, Br, Cl, F, OTs, OMs, OTf).