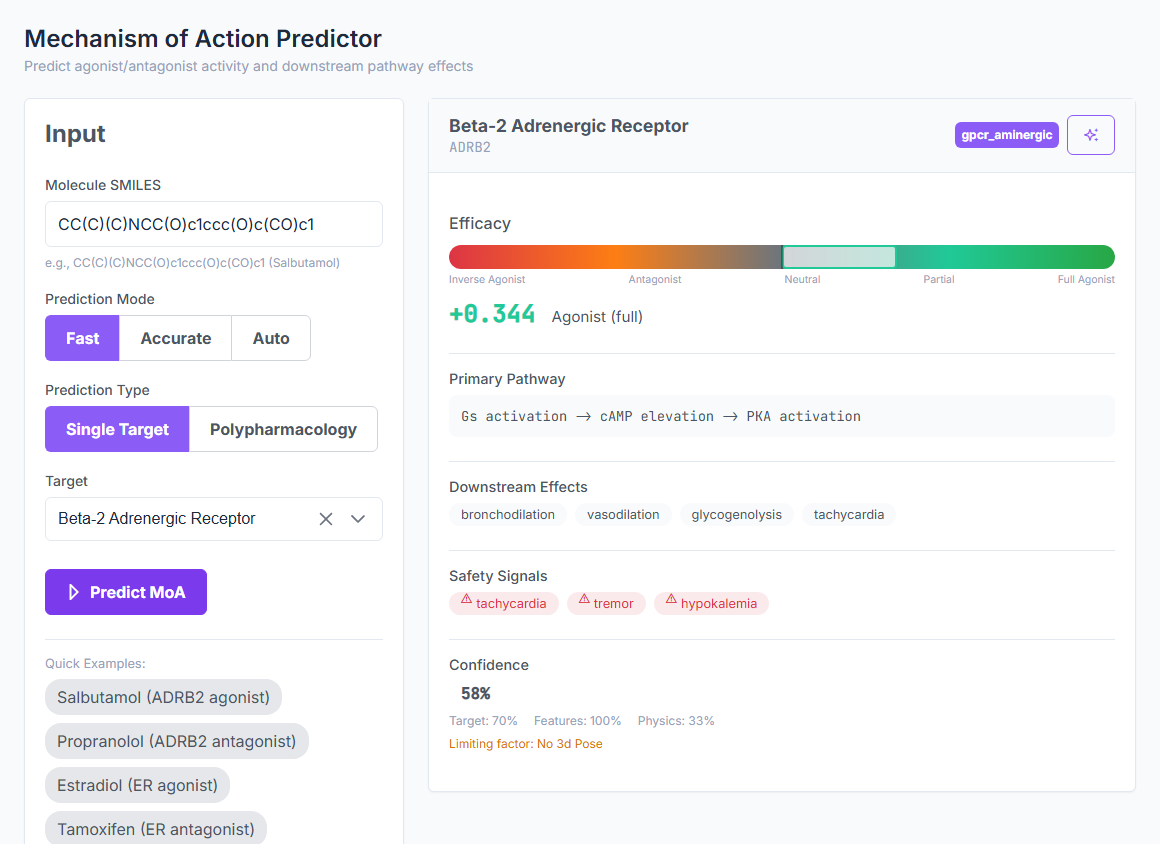
Binding affinity, off-target, microbiome,
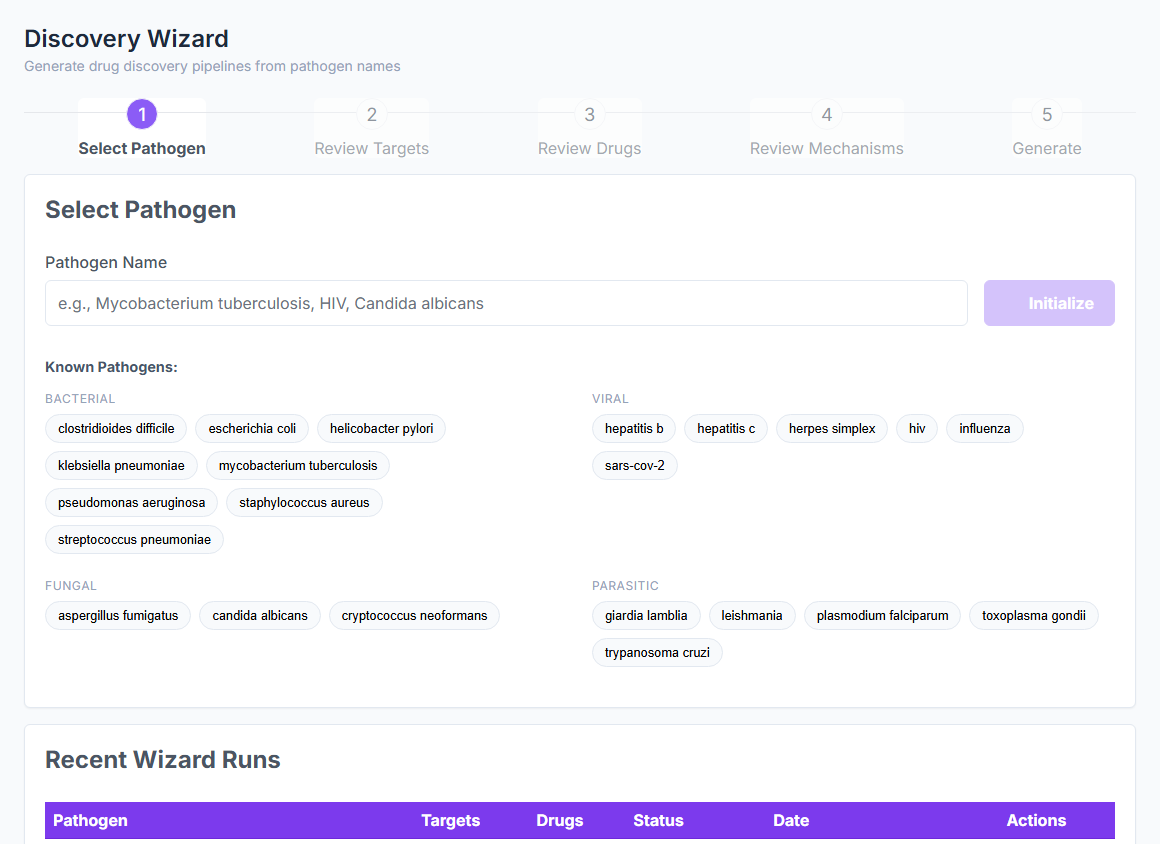
from a SMILES and a target name
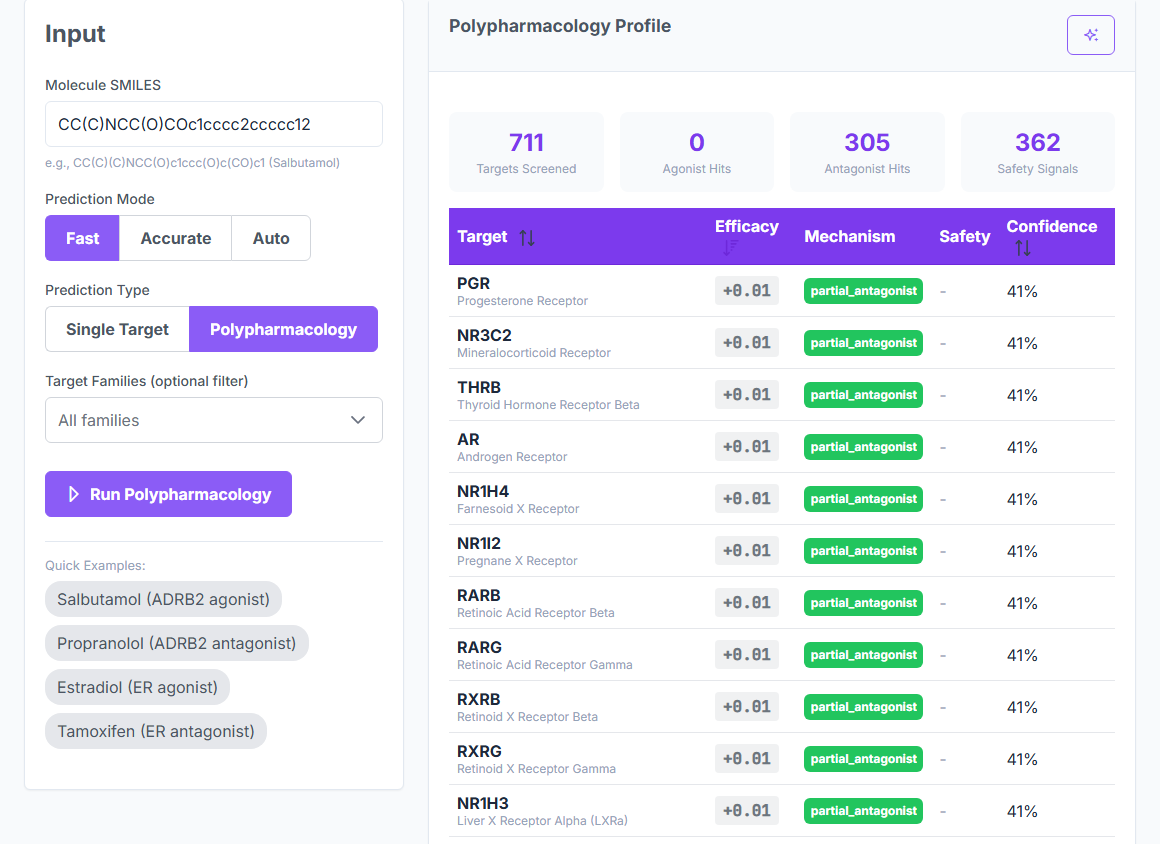
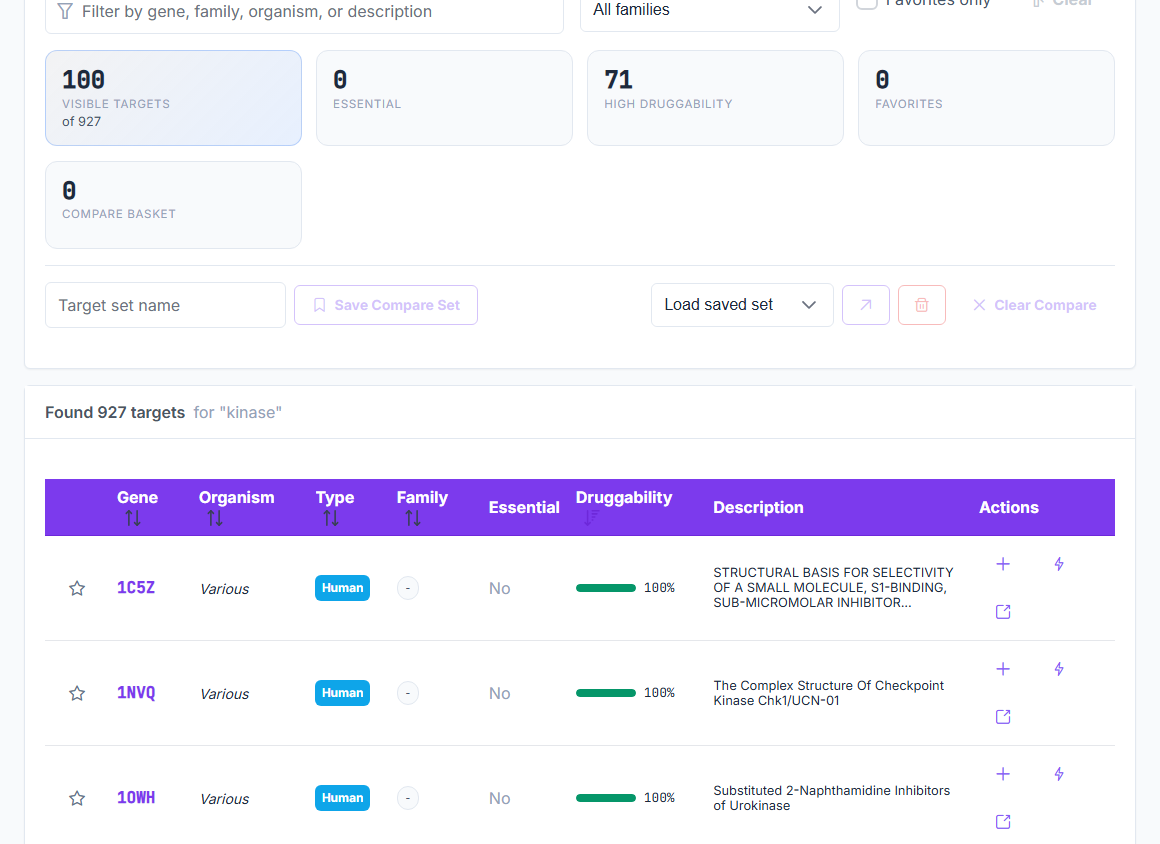
BioTarget scores pathogen efficacy, human off-target risk, and microbiome impact across 10,065 curated targets in 5 biological kingdoms. CASF-2016 validated. No crystal structure required. No training set. The whole multi-panel screen runs at 5,000+ predictions per second.