Full ADMET panel,
with confidence you can read
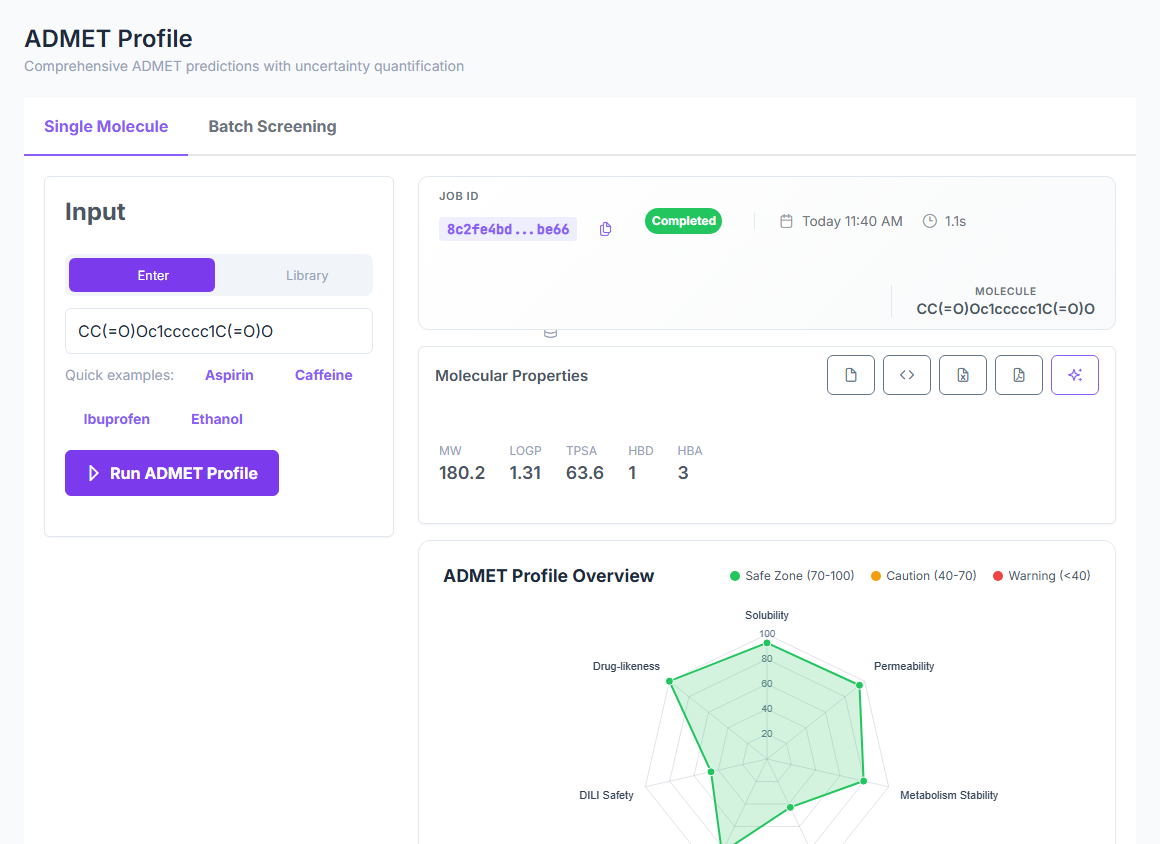
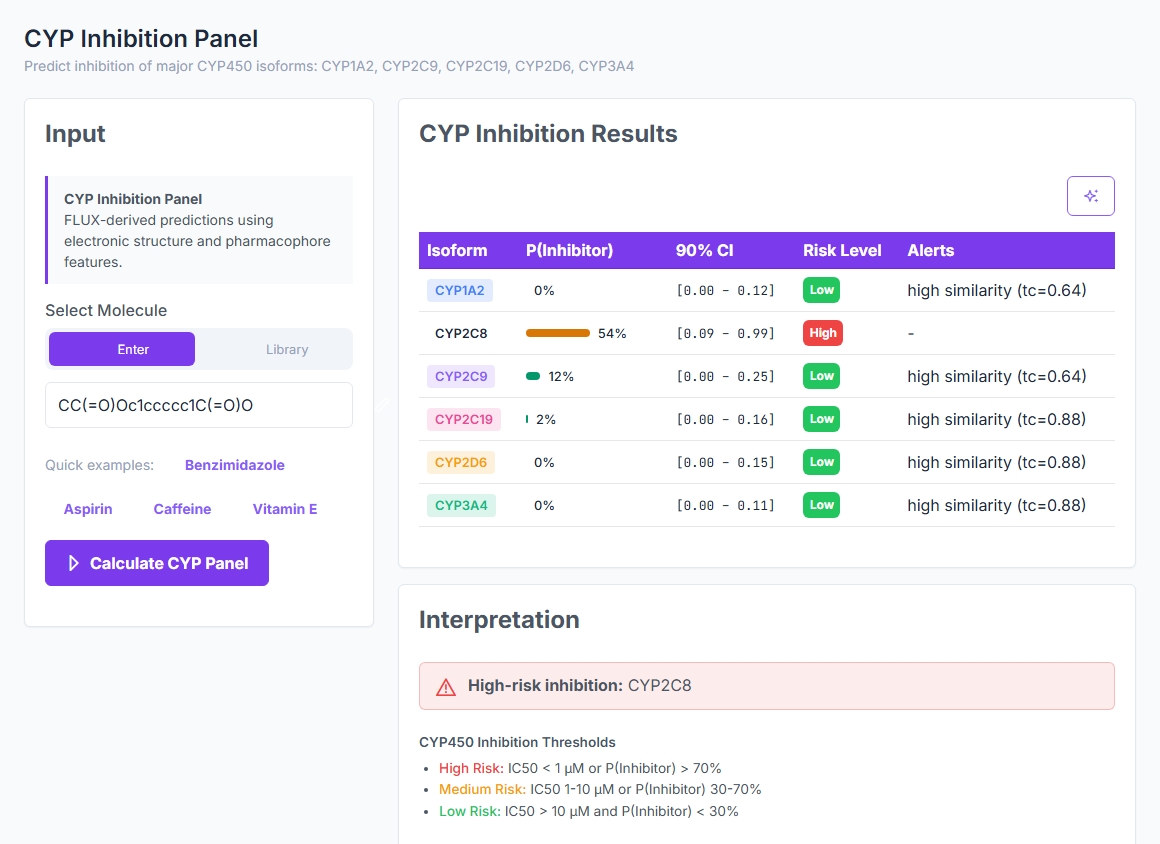
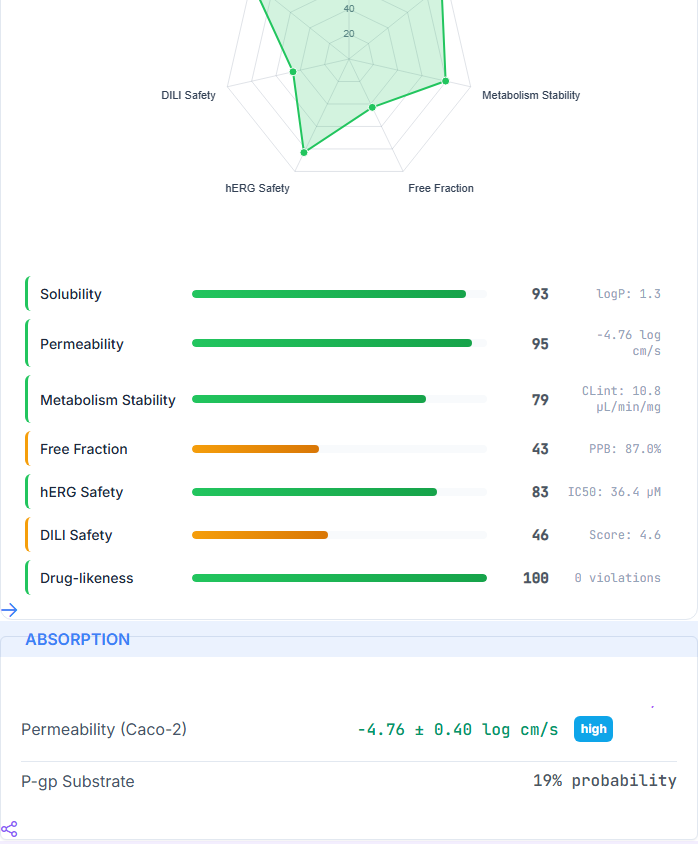
BBB, solubility, PPB, permeability, metabolism, hERG, DILI, and the full CYP panel — in one deterministic
framework. Every prediction comes with a calibrated confidence tier, so you know which numbers to trust and
which to verify. 5 of 8 endpoints sit at public-benchmark SOTA from pure physics —
Solubility, Metabolism, and PPB strict #1 on the TDC leaderboard; DILI AUROC 0.9597 on the comparable TDC
binary task; Caco-2 MAE 0.277 on the TDC caco2_wang scaffold-split test, matching the public
reference SOTA at 0.276 with zero training labels.
ADMET is the validated output spine of the larger Flux Pharmacology cascade — the same engine also produces binding affinity, selectivity, and repurposing endpoints from the same physical model.